OBJECTIVES
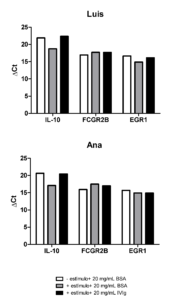
– To test whether stimulated B cells differentially express EGR1 (as in the anergy paper by Seité et al.) IL-10 and/or Fcgr2b genes upon IVIg treatment.
MATERIALS
– High Capacity RNA-to-cDNA Kit (Ref:4387406, Applied Biosystems)
– TaqMan Universal Master Mix II with UNG (Ref:4440042, Applied Biosystems)
– Human IL-10 Assay Hs00961622-m1 (Applied Biosystems)
– Human Fcgr2b Assay Hs01634996-s1 (Applied Biosystems)
– Human EGR1 Assay Hs00152928_m1 (Applied Biosystems)
– Human 18S Assay Hs99999901-s1 (Applied Biosystems)
PROTOCOLS
A) RT (20161117)
Samples were retrotranscribed according to the following protocol. A blank sample (w/o RNA template) as a negative control was added to the reaction.
| n=1 | MM (n=6+3=9) | |
|---|---|---|
| 2X RT Buffer | 10 mcL | 90 mcL |
| 20X RT Enzyme mix | 1 mcL | 9 mcL |
| Nuclease-free H2O | qsp. 20 mcL | - |
| Sample | up to 9 mcL* | - |
| Total per reaction | 20 mcL | Add 11 mcL per microtube |
100 ng of sample RNA were used in each 20 mcLreaction. The following volumes were added per microtube.
| Datos | ESTÍMULO | TTO | Cuantificación RNA | Volumen muestra RT (100 ng) | Volumen H2O RT (qsp 9 mcl) | ||
|---|---|---|---|---|---|---|---|
| Nº individuo | Nombre | ODN +anti-G/M/A | IVIg | Concentración (ng/mcL) | mcL | mcL | |
| 1 | LUIS | NO | NO | 13.8 | 7.2463768115942 | 1.7536231884058 | |
| SI | NO | 53.5 | 1.8691588785047 | 7.1308411214953 | |||
| SI | SI | 32.7 | 3.0581039755352 | 5.9418960244648 | |||
| 2 | ANA | NO | NO | 20.6 | 4.8543689320388 | 4.1456310679612 | |
| SI | NO | 145.6 | 0.68681318681319 | 8.3131868131868 | |||
| SI | SI | 76.1 | 1.3140604467806 | 7.6859395532194 |
The thermal cycler conditions were as follows, as indicated by Applied (total volume 20 mcL):**
| Step 1 | Step 2 | Step 3 | |
|---|---|---|---|
| Temperature (º C) | 37 | 95 | 4 |
| Time | 60 min | 5 min | Infinite Hold |
** Note: power went off at the first step of the RT protocol but sample integrity was preserved as proved by the 18S amplification seen in the 20161122 test qPCR.
C) qPCR (20161123)
Samples were analized in triplets and two blank samples were added to the reaction as negative controls: one sample w/o cDNA template (qPCR blank) and the RT negative control.
The following protocol was used:
| N=1 | MM per detector (n=18+3) | |
|---|---|---|
| Taqman Universal Master Mix II | 10 mcL | 210 mcL |
| Taqman Gene Expression Assay | 1 mcL | 21 mcL |
| cDNA template+ H2O | up to 9 mcL* | - |
| Total volume | 20 mcL | Add 11 mcL per well |
1,2 mcL of sample (100ng/20 mcL*1,2 mcL=6 ng) and 7,8 mcL of water were added per well. Since 12 wells (n=12+2=14) were analized per sample, a mastermix of 16,8 mcl of sample+ 109,2 mcl of water.
The thermal cycler conditions were as follows, as indicated by Applied (total volume 20 mcL):
| UNG incubation (Hold) | Polymerase activation (Hold) | PCR (40 cycles) | |
|---|---|---|---|
| Temperature (ºC) | 50 | 95 | 95 + 60 |
| Time (mm:ss) | 2:00 | 10:00 | 00:15 + 1:00 |
RESULTS